Electronic Journal of Academic and Special Librarianship
v.6 no.1-2 (Summer 2005)

Electronic Journal of Academic and Special Librarianshipv.6 no.1-2 (Summer 2005) |
 |
John M. Weiner
weiner@buffnet.net
Index terms are an important component in considering a scientific topic. In a real sense, the indexing terms represent the vocabulary and language of the topic. Study of these critical terms has employed human and machine techniques. Computerized indexing systems can accurately and completely recognize terms, but the different strategies for organizing and evaluating the concepts (i.e., informative terms) and related issues may not be effective in accomplishing the desired descriptive actions. This paper explored the results of two computer supported approaches in indexing scientific documents against a background of simple random generation of informative terms in varying sized text blocks. One method (RefViz) is based on the Latent Semantic Indexing with Multidimensional Scaling (LSI,MDS) approach. This system identifies potential indexing terms from the natural language text. The terms are stripped from the text, organized using statistical criteria and are used to classify documents based on assigned 'meanings' or themes. These themes are based on frequency and correlation and each document is assigned one. The second (Idea Analysis) also uses the natural language text. In this system, the terms are identified within the authors' sentences and couplets extracted. A single document may be represented by 100 or more term couplets representing the authors' thoughts. Both systems are superior to random sampling. The results suggested that indexing based on the authors' thoughts may be better than indexing based on statistical criteria.
The purpose of this report is to assess the value of different methods for computerized coding of informative terms and the implications associated with variations observed. Index terms and organizations of these terms have evolved into languages and complex arrangements depicting content and meaning associated with a scientific topic. [Brown 2000, Funk 1983, UMLS 2004] In addition to the traditional triage and retrieval of documents for manual processing, the newer computerized indexing systems tend to offer understanding of the topic by considering themes representing clusters of documents in the set. [Jenuwine 2004, Johnson 1993, Joubert 1993, Kashyap 2003] Depending on the system, a single document may be represented one or many times in the different clusters. Two approaches in computerized text mining (i.e., identification, extraction and organization of informative terms) were considered in this study. The systems differ in process. One seeks to determine themes using informative terms from the text and developed these themes, based on term triads, organized using statistical criteria. That system is latent semantic indexing with multiple dimensional scaling (LSI,MDS). [Dumais 1996, Foltz 1998, Landauer 1998] The other identifies author-provided couplets of informative terms within sentences. That system is author semantic indexing (ASI). [Berman 2004, Weiner 1983, Weiner 2004]
Latent Semantic Indexing: Latent Semantic Indexing (LSI) is a modern, sophisticated text analysis approach intended to identify and display the themes predominant in a set of documents. [Dumais 1996, Landauer 1994, Landauer 1998, Rehder 1998, Yu 2002] The system is used in computerized text mining and is currently thought to be effective in describing conceptual themes inherent in the document set analyzed. The informative terms are identified and extracted from the text and organized as a list. A matrix is constructed composed of the documents on the horizontal axis and the informative terms on the vertical axis. The terms in a particular document are noted in the column as a set of frequencies. The terms that are absent are identified as zeroes.
By rearranging the vectors, those with the largest number of similar terms can be placed adjacent to one another. The ones with the larger number of dissimilar terms can be separated from the rest.[Foltz 1998] To accelerate the computations, similarity scores could be calculated using different models. One model is multidimensional scaling (MDS).[Yu 2002] It is designed to identify three key terms that 'adequately' describe the vector for the document. These summary vectors are arranged until the similarity scores for adjacent neighbors are highest and clusters of documents are formed representing the different themes in the document set.
These clusters can be displayed graphically. A set of documents containing terms that are similar would form a graph with the clusters arranged in a circle. As dissimilarity is introduced, the circular shape changes to an elliptical one. A set of documents with highly dissimilar clusters would have an elongated elliptical appearance.
Figure 1. Example of Galaxy Display showing Specific Cluster of Documents using
RefViz Software.

Figure 1 illustrates the LSI with MDS approach using a software product entitled RefViz. [RefViz 2003] This is a data visualization and analysis software from the makers of EndNote, ProCite, and Reference Manager. The creators described their product as "… an essential tool designed to help researchers evaluate references easily, plan future projects and publish their work. RefViz provides users with a powerful way to explore reference literature visually. It analyzes large numbers of references by thematic content and presents an at-a-glance overview of the main topics discussed in the reference set." References are organized visually using the Galaxy graphic, a proximity map showing clusters of articles based on thematic context. References or groups of references containing common themes are placed in proximity to one another. The resulting graphic shows a logical flow of one concept to another across the Galaxy.
This approach seeks to build a new type of text warehouse containing themes defined using statistical criteria, document grouping identification, and copies of the original documents. With this resource, the terms important in identifying the cluster of documents are known in advance and they can be retrieved based on that prior knowledge. Figure 1 showed an example of this output. The full set of documents entered into the text warehouse are grouped according to combinations of informative terms and displayed graphically. By highlighting a particular cluster, the terms involved in defining the cluster are shown together with access to the documents satisfying the statistical criteria. As seen, Group 22 contains 74 references and these are characterized by the triad consisting of p53, breast, and melanoma. The documents described epidemiologic research.
Author Semantic Indexing: Author Semantic Indexing is based on analysis of the concepts and relationships described in the philosophy of information.[Fallis 2004, Floridi 1999, Furner 2004, Goldman 2001, Hernon 1995] Figure 2 summarizes the perceptions regarding information. Links shown are those involving pairs of the terms as provided by the authors in their sentences. Of the 40 terms suggested as representing information and related concepts, 10 have particular interest (shown with black background and white text) in terms of justifying a computerized process.
Figure 2. Information and Related Concepts.*

*From Analysis of Information Philosophy Literature. Map subject to change.
As seen in Figure 2, information has been linked with cognition, communication, creative, knowledge, meaning, and relevance. Knowledge was linked with cognition, creative, information, interpretation, mental act, public, and truth. Each of these terms implies subjective criteria varying in specificity from person to person. Further, the definitions of these terms do not contain those that would motivate development of a system. The figure shows another cluster of terms linked to meaning. The links involve meaning with interpretation, informativeness, language and word. Interpretation is linked with sentence, thought, creative and information. Understanding is linked with language and language is linked with thought and sentence. This restricted swarm of concepts is of interest in considering computerized text analysis. The philosophers linked meaning and word but did not relate word to sentence or thought. The existing links involving interpretation, sentence, thought and creative and those involving language, sentence and thought suggest an approach that could be accomplished using computer methods. The common elements in these swarms are sentence and thought. While word plays an important role in current computerized text analysis, the information philosophers have not linked that term with sentence, thought or creative.
Random Occurrence of Informative Terms: A simulation study [Bruce 2000, Weiner 2005] was performed to determine the likely effects of text length on recognition of informative term couplets and triads. A set of hypothetical abstracts was considered with each abstract composed of sentences of varying in length from 4 to 512 words. There were 10 sentences to each abstract. The text was composed of numbers representing informative terms. Pairs or triads of these terms were assigned meaning and similarity scores computed. The vocabulary was represented by numbers between 1 and 3000. Similarity was scored by determining the duplicates and triplicates found in the randomly generated strings of 'words' tested. A document set consisted of 1000 abstracts.
Figure 3. Frequency of All Duplicates in 1000 Abstracts, each with 10 Sentences.
The number of words in a sentence varied from 4 to 512.

Figure 3 showed the relative frequency of all duplicates in sentences containing 4 to 512 terms. As seen, the occurrence of pairs was more than 90% when the length of the text string analyzed consisted of 8 to 20 words. The typical abstract of 250 words has a probability of recognizing random words in couplet patterns approximated 20% of the time. Triads in random word sentences occurred approximately 70 percent of the time when the sentences contained 8-10 words. In abstracts of approximately 250 words, the occurrence of triads was 5% - 10%.
These simulation results set the stage for the comparison described in this report. The RefViz method identifies informative terms using the entire abstract as one text block. The Idea Analysis system identifies informative terms using each author's sentence. In that system, a term must be linked with another one within the sentence to be included in the vocabulary.
RefViz [RefViz 2003] and Idea Analysis [Weiner 1981] software systems were compared using the epidemiologic research reports entered into Medline. The two computerized systems were used to build new repositories consisting of the vocabularies and related data representing the respective system's definition of a text warehouse. Epidemiologic research was selected because of the diverse topics considered as well as the employment of multidisciplinary teams. The two system vocabularies were used to compare with the one used by subject experts. The experts' descriptions were retrieved from the 2004 entries in CRISP (Computer Retrieval of Information on Scientific Projects). [CRISP 2004] This study was a step in attempting to provide data dealing with issues of agreement between a credible gold standard and computerized text mining systems.
One hundred twenty-eight thousand, one hundred and forty (128,140) articles representing epidemiologic research entered into Medline during 2000 – 2003 were analyzed. These reports were processed using RefViz and Idea Analysis. The time required to prepare the two resources were respectively, 0.05 and 0.06 minutes per article. This analysis was designed to compare the degree of agreement between the experts' vocabulary and those developed using the text mining systems.
A second analysis considered different topics to estimate the stability of the agreement between Idea Analysis and the experts' vocabularies. This analysis considered 1,608 articles entered into Medline for 2004. The topics included were: emergency medicine (254), internal medicine (879), aging and exercise (161), family medicine (111), and neuropsychiatry (183). Random samples of 15 reports were retrieved in each of these subjects from CRISP and the vocabularies compared.
Figure 4 shows the results for the 20 randomly selected documents from the 4218 CRISP epidemiologic research projects for 2004 plus 11 additional randomly selected reports dealing with respectively, asbestos or asthma research in the same epidemiology set. The identity line represents equal percent agreement for each system. There was only one report where both systems agreed and that one involved 5 commonly used terms. The remaining random abstracts showed consistently fewer matches for RefViz than for Idea Analysis. The median percent agreement for RefViz was 28% and for Idea Analysis, 80%.
Figure 4. Percent Agreement of Idea Analysis and RefViz with CRISP
Approved Research Project Abstracts.
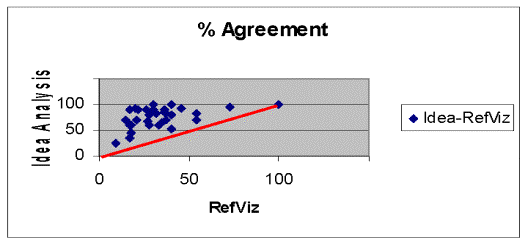
Table 1 shows the matches between CRISP and vocabularies built using Idea Analysis. The subjects studied were Emergency Medicine (81% agreement), Internal Medicine (91% agreement), Aging and Exercise (69%), Family Medicine (59%), and Neuropsychiatry (76%). The median percent agreement was 76%.
Table 1. Probability of Matching Vocabularies
Using CRISP and Idea Analysis Knowledge Base Analyses.
| Emergency Medicine | 81 |
| Internal Medicine | 91 |
| Aging & Exercise | 69 |
| Family Medicine | 59 |
| Neuropsychiatry | 76 |
This report compared experts' use of informative terms against two computerized text mining approaches. The first, latent semantic indexing with multidimensional scaling (LSI,MDS), is based on sophisticated statistical procedures. [Landauer 1994, Rehder 1998, Shatkay 2003, Yang 1991, Yu 2002] The second, author semantic indexing (ASI), is based on the premise that authors convey their thoughts within sentences. [Chen 1988, Hoffman 1980, Piniewski-Bond 2001, Weiner 2004] The differences observed, using the epidemiologic research reports, showed that the ASI approach was superior to LSI,MDS. However, both computerized systems were superior to random generation of informative term couplets or triads, considering the typical 250 word abstract.
In addition, the comparison of expert vocabularies (from CRISP) with different medical topics using Idea Analysis showed that the estimate of agreement from the epidemiologic literature (81%) was close to the estimate from the diverse topics (76%).
The assumption supporting analysis of the complete text block as an entity is that the author organized the text to convey a set of related thoughts. As such, arbitrary linking of terms across sentences, paragraphs or pages is appropriate in discerning the relationships envisioned by the author. [Auerbuch 2004, Brown 2000, Friedman 2004, Hirschman 2002, Pennebaker 2004, Zoo 2003] That assumption is challenged by these data as well as previous reports.[Sigurd 2004, Sparck Jones 1972] These studies suggest that computerized systems focusing on the contents within authors' sentences may develop a vocabulary describing the topic that is more in agreement with the one employed by the subject experts. This obvious finding contradicts the approach employed in the majority of computerized systems. In those, the software employs larger blocks of text and uses statistical criteria (frequency, correlation or both) to select terms and assign meaning to documents.
The agreement between experts' vocabularies and Idea Analysis is interesting in that this study considered the matching given a particular topic rather than the more realistic multiple topic selection process used by authors. Single focus is a major limitation in the present approach but important in clarifying the role of a software system in text mining. Investigators may pick and choose concepts and relationships from other subjects and introduce them into their research within a given topic. [Smalheiser 1998, Swanson 1990, Weeber 2003] As such, the vocabulary used in an approved research proposal (in CRISP) need not agree completely with the vocabulary in the literature. However, the agreement should be better than random generation. In addition, the conditions employed in the system providing the highest agreement should be further considered. The array of conceptual issues within a topic would not be expected to include those from some other topic unless previously introduced by investigators and recorded in the topic literature.
Contextual importance must be considered. [Brown 2000, Leroy 2001, Sparck Jones 1972] Systems, using frequency as a criterion for recognizing important terms, cannot differentiate between those that are contextually relevant versus those that are not. While all of the computerized approaches use the authors' terms as the basis of the analysis, ASI uses the authors' combinations of terms and restricts the development of vocabulary to only those terms found as elements in authors' thoughts. Other systems form statistical combinations of terms, thus minimizing the meaning assigned by the authors.
The results show the feasibility of comparing authors' text (from CRISP) and the vocabulary constructed using Idea Analysis from analysis of the Medline reports. The software can recognize sentences, informative terms, and term couplets in each sentence. CRISP documents can be retrieved and processed using manual procedures to identify the informative terms in that text. While these analytic approaches are different, the resulting data can be compared and conclusions reached regarding the feasibility of computerized text mining in analysis of scientific literature.
This study compared vocabularies generated by computerized text mining systems with that employed by subject specialists. This comparison is important in that indexing terms have taken on importance beyond that expected in simple identification and retrieval. Indexing terms represent a language describing the topic. The terms may be arranged to depict the inherent concepts and relationships comprising the topic. The study showed that the two text mining systems considered were superior to simple random generation of informative term couplets or triads. The LSI,MDS system is based on a sophisticated statistical model. That system developed a vocabulary for the epidemiologic literature that matched the experts approximately 30% of the time. The ASI system, based on information philosophy concepts, matched the experts approximately 80% of the time. The question suggested by these analyses is, "What conditions should be stressed in building a computerized indexing system – arbitrary assignment of importance based on statistical measures or subject expert assignment of importance based on knowledge and experience?"
1. Auerbuch M, Karson TH, Ben-Ami B, Maimon O, Rokach L. Context sensitive Medical Information Retrieval. Medinfo. 2004;2004:282-6.
2. Berman JJ. Doublet method for very fast autocoding. BMC Med Inform Decis Mak. 2004 Sep 15;4(1):16.
3. Brown PJ, Sonksen P. Evaluation of the quality of information retrieval of clinical findings from a computerized patient database using a semantic terminological model. J Am Med Inform Assoc. 2000 Jul-Aug;7(4):392-403.
4. Bruce P, Simon JL, Oswald T. Resampling Stats. Arlington, VA, 2000. Available at: http://www.stats@resample.com. Accessed May 16, 2005.
5. Chen J. The natural structure of scientific knowledge: an attempt to map a knowledge structure. Journal of Information Science. 1988;14:131-139.
6. CRISP. Available at: http://crisp.cit.nih.gov/. Accessed May 16, 2005.
7. Dumais ST, Landauer TK, Littman ML. Automatic cross-linguistic information retrieval using Latent Semantic Indexing. In: SIGIR'96 - Workshop on Cross- Linguistic Information Retrieval. 1996:16-23.
8. Fallis D. On Verifying the Accuracy of Information: Philosophical Perspectives. Library Trends. 2004;52:427-446.
9. Floridi L. Philosophy and Computing: An Introduction. London: Routledge; 1999.
10. Foltz PW, Kintsch W, Landauer TK. The Measurement of Textual Coherence with Latent Semantic Analysis. Discourse Processes. 1998;25:285-307.
11. Friedman C, Shagina L, Lussier Y, Hripcsak G. Automated encoding of clinical documents based on natural language processing. J Am Med Inform Assoc. 2004 Sep- Oct;11(5):392-402.
12. Funk ME, Reid CA. Indexing consistency in MEDLINE. Bull Med Libr Assoc. 1983 Apr;71(2):176-83.
13. Furner J. Information Studies Without Information. Library Trends. 2004; 52(3):463-487.
14. Goldman AI. Experts: Which ones should you trust? Philosophy and Phenomenological Research. 2001;63(1):85-110.
15. Hirschman L, Park JC, Tsujii J, Wong L, Wu CH. Accomplishments and challenges in literature data mining for biology. Bioinformatics. 2002;18(12):1553-61.
16. Hoffman E. Defining information: an analysis of the information content of documents. Information Processing and Management 1980;16:291-304.
17. Jenuwine ES, Floyd JA. Comparison of Medical Subject Headings and text-word searches in MEDLINE to retrieve studies on sleep in healthy individuals. J Med Libr Assoc. 2004 Jul;92(3):349-53.
18. Johnson SB, Aguirre A, Peng P, Cimino J. Interpreting natural language queries using the UMLS. Proc Annu Symp Comput Appl Med Care. 1993:294-8.
19. Joubert M, Fieschi M, Robert JJ. A conceptual model for information retrieval with UMLS. Proc Annu Symp Comput Appl Med Care. 1993:715-9.
20. Kashyap V. The UMLS Semantic Network and the Semantic Web. AMIA Annu Symp Proc. 2003:351-5.
21. Landauer TK, Dumais ST. Latent semantic analysis and the measurement of knowledge. In: Kaplan RM, Burstein JC, eds. Educational testing service conference on natural language processing techniques and technology in assessment and education. Princeton, NJ: Educational Testing Service;1994.
22. Landauer TK, Foltz PW, Laham D. Introduction to Latent Semantic Analysis. Discourse Processes. 1998;25:259-284.
23. Leroy G, Chen H. Meeting medical terminology needs--the Ontology-Enhanced Medical Concept Mapper. IEEE Trans Inf Technol Biomed. 2001 Dec;5(4):261-70.
24. Pennebaker J. What our words say about us: Use of computerized text analysis in research and practice. In: International Society for Quality of Life Research, ISOQOL 2004 Symposium; June 27-29, 2004.
25. Piniewski-Bond JF, Buck GM, Horowitz RS, Schuster JH, Weed DL, Weiner JM. Comparison of information processing technologies. J Am Med Inform Assoc. 2001 Mar-Apr;8(2):174-84.
26. RefViz. Available at: http://www.RefViz.com. Accessed May 16, 2005.
27. Rehder B, Schreiner ME, Wolfe MB, Laham D, Landauer TK, Kintsch W. Using Latent Semantic Analysis to assess knowledge: Some technical considerations. Discourse Processes. 1998;25:337-354.
28. Shatkay H, Feldman R. Mining the Biomedical Literature in the Genomic Era: An Overview. J Comput Biol. 2003;10(6):821-855.
29. Sigurd B, Eeg-Olofsson M, van Weijer J. Word length, sentence length and frequency. Zipf revisited. Studia linguistica. 2004;58:37-52.
30. Smalheiser NR, Swanson DR Using ARROWSMITH: a computer-assisted approach to formulating and assessing scientific hypotheses. Comput Methods Programs Biomed. 1998 Nov;57(3):149-53.
31. Sparck Jones K. A statistical interpretation of term specificity and its application in retrieval. Journal of Documentation. 1972;28(1):11—21.
32. Swanson DR. Medical literature as a potential source of new knowledge. Bull Med Libr Assoc. 1990 Jan;78(1):29-37.
33. Unified Medical Language System. Available at: http://www.nlm.nih.gov/pubs/factsheets/umls.html. Accessed May 16, 2005.
34. Weeber M, Vos R, Klein H, De Jong-Van Den Berg LT, Aronson AR, Molema G. Generating hypotheses by discovering implicit associations in the literature: a case report of a search for new potential therapeutic uses for thalidomide. J Am Med Inform Assoc. 2003 May-Jun;10(3):252-9.
35. Weiner JM, Shirley S, Gilman NJ, Stowe SM, Wolf RM. Access to data and the information explosion: oral contraceptives and risk of cancer. Contraception. 1981 Sep;24(3):301-13.
36. Weiner JM: Text Analysis and Basic Concept Structures. Information Processing and Management. 1932;19:313-319.
37. Weiner JM, Schuster JHR, Horowitz RS, McAfoos WP, Piniewski-Bond J. Fantasies in Processing. Baltimore: American Literary Press; 2004.
38. Weiner, JM. Testing Conditions in Latent Semantic Indexing. Submitted for Publication 2005.
39. Yang Y, Chute CG. A schematic analysis of the Unified Medical Language Sy stem. Proc Annu Symp Comput Appl Med Care. 1991;204-8.
40. Yu C, Quadrado J, Ceglowski M, Payne JS. Patterns in Unstructured Data. Available at: http://javelina.cet.middlebury.edu/lsa/out/cover_page.htm. Accessed May 16, 2005.
41. Zou Q, Chu WW, Morioka C, Leazer GH, Kangarloo H. IndexFinder: a method of extracting key concepts from clinical texts for indexing. AMIA Annu Symp Proc. 2003;763-7.